Assignment 3.3 Guide
In Assignment 3.3 you will conduct basic sequence and structural analysis, as well as 3D modeling of your protein of interest that you selected in Assignement 3.2. When a scientist begins working closely with one protein or a family of proteins she first gathers all the basic information about the protein(s). You will do the same for your protein of interest. The ExPASy website has a wide range of free software tools listed under "Categories --> proteomics --> protein structure" that will be of great use for this Assignment and future characerizations of proteins. Below you find the required data you will obtain for this assignment and a list of selected tools that will help you along your way. Some information may also be best obtained from Phyre2, described below.
DATA to Compile:
- Length of protein (number of amino acids)
- Predicted Molecular weight
- Predicted pI (Isoelectric point)
- Most abundant amino acid
- Extinction coefficient
- How many helices are predicted
- How many Beta strands (sheets)
Selected online tools:
To make predictions about drug design and protein-protein interactions it helps to have a 3D model of your protein of interests. There are many high-end commercial programs that will create highly accurrate models, and there are freely available tools that are more than sufficient for preliminary analysis of proteins. One such freely available tool is called Phyre2 and can be found here: http://www.sbg.bio.ic.ac.uk/phyre2/html/page.cgi?id=index. This program takes several hours to run so plan ahead and do not leave it until the last minute. To submit your sequence for analysis you must enter your email address, it is recommended you enter a job description and finally you select Phyre Search.

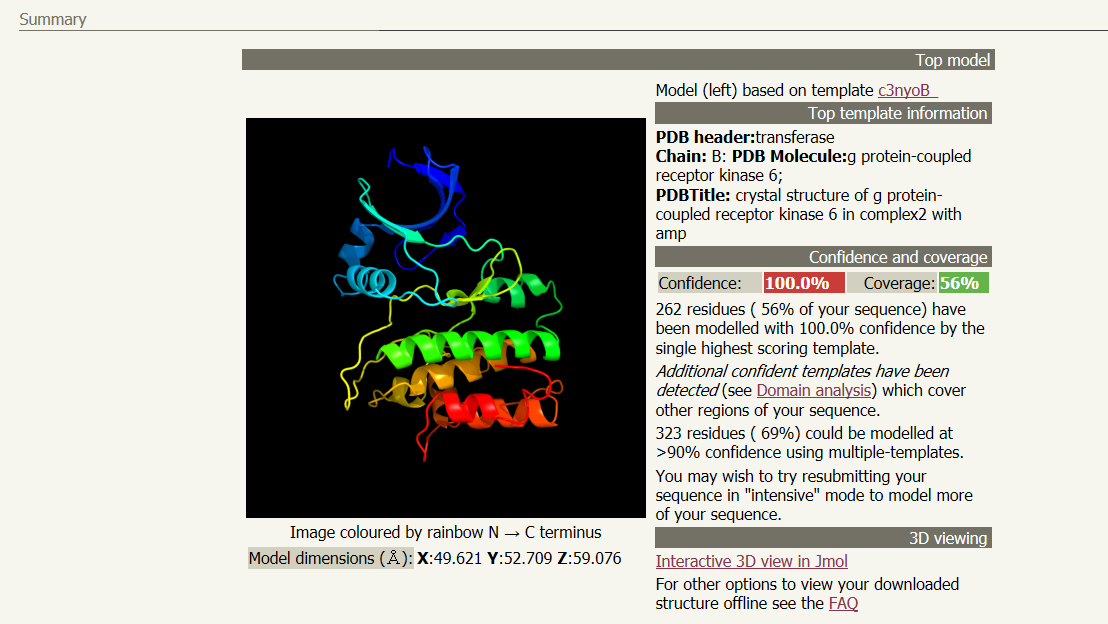
A link will be emailed to you after several hours with the results of the analysis. Follow the link given in the email to your results page. You will see a ribbon style predicted structure for your protein. Other information, including what previously determined structure was used as model and how confident the program is with the predicted stucture can be found next to the structure. In the email will also be a .pdb file containing the data required for other software programs to visualize your 3D structure. A suggested list the viewers and visualizers is below.
- Swiss PDB Viewer "DeepView" (You will need to download the program and run it locally)
- Protein Picture Generator
The next section in your results file is the "Secondary structure and disorder prediction", where you are show the number and location of disordered regions, alpha helices, and beta strands (sheets). The sections below secondary strucure analysis detail domain predictions, binding site predictions, and transmembrane helix predictions. You should explore the wealth of data provided by this one tool. Using the skills learned in Unit 1, create a presentation that introduces and describes the characteristics of your selected protein. You are required to create a table detailing the information gathered from the various analysis tools you used. In your presentation be sure to note the tools used to aquire your data.

